|
rbmatlab
1.16.09
|
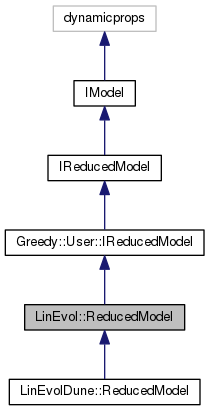
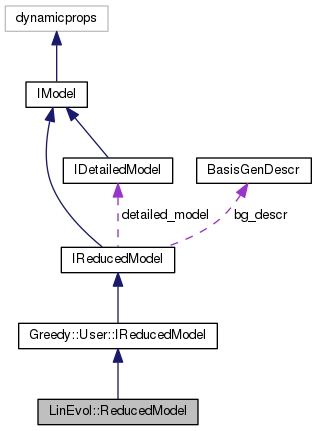
reduced model for linear evolution problems as given by a LinEvol.DetailedModel.
This is compatible with Greedy.User.IReducedModel and can therefore make use of detailed data objects created by Greedy algorithms.
Definition at line 18 of file ReducedModel.m.
Public Member Functions | |
| ReducedModel (LinEvol.DetailedModel dmodel,BasisGenDescr bg_descr) | |
| Constructor for the reduced model. More... | |
| function a0 = | rb_init_values (Greedy.DataTree.Detailed.RBLeafNode detailed_data, decomp_mode) |
| function computing initial values for a reduced simulation. More... | |
| function
rb_sim_data = | rb_simulation_impl (LinEvol.ReducedData reduced_data) |
| function, which performs a reduced basis online simulation for the parameter vector \(\mu \in {\cal P} \subset \mathbb{R}^p\), which is assumed to be set by IDetailedModel.set_mu() More... | |
| function
rb_sim_data = | rb_reconstruction (LinEvol.DetailedData detailed_data, rb_sim_data) |
| (trivial) function computing a detailed reconstruction by linear combination of the coefficients in the simulation data with the orthonormal reduced basis RB More... | |
| function LinEvol.ReducedModel c = | copy () |
| function that deep copies this handle class More... | |
 Public Member Functions inherited from Greedy.User.IReducedModel Public Member Functions inherited from Greedy.User.IReducedModel | |
| IReducedModel (IDetailedModel dmodel,BasisGenDescr bg_descr) | |
| constructor for this reduced model interface More... | |
| function
Greedy.User.ReducedData reduced_data = | gen_reduced_data (Greedy.User.IDetailedData detailed_data) |
Constructs the reduced_data object holding low dimensional data needed for efficient reduced simulations with rb_simulation(). More... | |
| function
rb_sim_data = | rb_simulation (Greedy.User.ReducedData reduced_data) |
| forwards the reduced simulation to the method rb_simulation_impl() after getting a suitable reduced data leaf element. More... | |
| virtual function
rb_sim_data = | rb_simulation_impl (Greedy.User.IReducedDataNode reduced_data) |
| implementation of the reduced simulation More... | |
 Public Member Functions inherited from IReducedModel Public Member Functions inherited from IReducedModel | |
| IReducedModel (IDetailedModel dmodel, bg_descr) | |
| Constructor of a reduced model. More... | |
| function iseq = | eq (IReducedModel other) |
Comparison operator checking whether the underlying detailed_model members of this and other are equal. More... | |
| function IReducedData reduced_data = | gen_reduced_data (detailed_data) |
Constructs the reduced_data object holding low dimensional data needed for efficient reduced simulations with rb_simulation(). More... | |
| function
reduced_data_subset = | extract_reduced_data_subset (IReducedData reduced_data) |
Extracts a subset of the reduced_data generated by gen_reduced_data(). More... | |
| virtual function
rb_sim_data = | rb_simulation (reduced_data) |
| Executes a reduced simulation and optionally an error estimation. More... | |
| virtual function
rb_sim_data = | rb_reconstruction (detailed_data, rb_sim_data) |
| reconstructs the reduced simulation snapshots generated by rb_simulation() in the reduced space \({\cal W}_{\text{red}}\). More... | |
| function U = | get_dofs_from_sim_data (sim_data) |
extracts the \(H\) dimensional Dof vector from the sim_data structure More... | |
| function IDetailedData detailed_data = | gen_detailed_data (model_data) |
| initiates the reduced basis generation process More... | |
| function p = | plot_sim_data (model_data, sim_data, plot_params) |
| plots the simulation data as returned by detailed_simulation() More... | |
| function
model_data = | gen_model_data () |
| generates large model data. More... | |
| function
sim_data = | detailed_simulation (model_data) |
| executes a detailed simulation for a given parameter More... | |
| function this = | set_mu (mu) |
| Sets the active parameter vector \(\mu \in {\cal M}\) used for simulations on this model. More... | |
| function mu = | get_mu () |
| returns the active parameter vector \(\mu \in { \cal M }\) More... | |
| function
rb_size = | get_rb_size (detailed_data) |
| returns the size of the generated reduced basis by the IDetailedData class. More... | |
| function | couple_N_and_M (detailed_data, ratio, factor) |
| sets all the basis sizes for reduced simulations by a ratio with respect to the maximum possible basis size and a factor between RB and EI basis sizes. More... | |
| function this = | set_Mratio (detailed_data, ratio) |
| in case of multiple operators subject to empirical interpolation, this sets number of reduced basis functions used for reduced simulations by specifying a ratio. More... | |
| function Mratio = | get_Mratio (detailed_data) |
| in case of multiple operators subject to empirical interpolation, this gets the mean of ratio between the number of reduced basis functions used for reduced simulations and the maximum possible. More... | |
| function
varargout = | subsref (S) |
| forwarding of fieldnames access to the underlying detailed_model description More... | |
Static Public Member Functions | |
| static function Delta = | get_estimators_from_sim_data (rb_sim_data) |
| static function Delta = | get_estimator_from_sim_data (rb_sim_data) |
| Static helper method returning an error estimator for the whole reduced trajectory \(\{u_{\text{red}}(\cdot, t^k)\}_{k=0}^{K}\) generated by rb_simulation(). More... | |
 Static Public Member Functions inherited from IModel Static Public Member Functions inherited from IModel | |
| static function ok = | struct_check (descr, checks) |
| executes checks on the fields of a structure object More... | |
Public Attributes | |
| enable_error_estimator = true | |
 Public Attributes inherited from IReducedModel Public Attributes inherited from IReducedModel | |
| descr | |
| The description structure holding information about the analytical parametrized problem and its discretization. More... | |
| decomp_mode | |
| mu_names | |
| cell array of strings describing the parameters of the model More... | |
| mu_ranges | |
| cell array of vectors of size two defining the allowed interval range for the parameter components More... | |
| verbose | |
| an integer defining the verbosity level of information output during basis generation More... | |
| debug | |
| an integer defining the debugging level controlling error output and extra tests during basis generation More... | |
| crb_enabled = false | |
| flag indicating whether this model depends on collateral reduced basis spaces. More... | |
| ::IDetailedModel | detailed_model |
| an object which shall be reduced | |
| ::BasisGenDescr | bg_descr |
| a structure defining the basis generation routines and data structures. | |
| enable_error_estimator | |
| boolean flag indicating whether during an rb_simulation() an a posteriori error estimator shall be computed. More... | |
| N = 0 | |
| control variable for the size of the reduced basis used for reduced simulations. By default this is equal to the size of the generated reduced basis. More... | |
| M = 0 | |
| control variable for the size of the (collateral) reduced basis used for empirical interpolations. By default this is equal to the size of the generated reduced basis. More... | |
| Mstrich = 0 | |
| control variable for the number of (collateral) reduced basis vectors used for error estimation. By default this is equal to zero. More... | |
 Public Attributes inherited from IModel Public Attributes inherited from IModel | |
| num_cpus = 4 | |
| The number of CPUs used for parallel sessions. More... | |
| decomp_mode | |
| Decomposition operation mode. More... | |
| mu_names | |
| cell array of strings describing the parameters of the model More... | |
| mu_ranges | |
| cell array of vectors of size two defining the allowed interval range for the parameter components More... | |
| verbose | |
| an integer defining the verbosity level of information output during basis generation More... | |
| debug | |
| an integer defining the debugging level controlling error output and extra tests during basis generation More... | |
Static Public Attributes | |
| static const | ddescr_checks |
| This constant is used for a consistency check of the model descr with help of IModel.struct_check() More... | |
 Static Public Attributes inherited from IModel Static Public Attributes inherited from IModel | |
| static const | time_checks |
| This constant can be used for a consistency check of time evolution members in the ModelDescr with help of IModel.struct_check() More... | |
| LinEvol.ReducedModel.ReducedModel | ( | LinEvol.DetailedModel | dmodel, |
| BasisGenDescr | bg_descr | ||
| ) |
Constructor for the reduced model.
| dmodel | object specifying how the high dimensional data can be computed. |
| bg_descr | structure specifying how the reduced basis shall be generated. |
descr — descr Definition at line 59 of file ReducedModel.m.
|
virtual |
function that deep copies this handle class
| c | an object which is a deep copy of this object. |
Implements IReducedModel.
Reimplemented in LinEvolDune.ReducedModel.
Definition at line 144 of file ReducedModel.m.

|
staticvirtual |
Static helper method returning an error estimator for the whole reduced trajectory \(\{u_{\text{red}}(\cdot, t^k)\}_{k=0}^{K}\) generated by rb_simulation().
| rb_sim_data | struct holding reduced simulation data returned by IReducedModel.rb_simulation() . struct holding reduced simulation data returned by IReducedModel.rb_simulation() . |
| Delta | This is a (K+1) x 1 vector of estimates \(\eta^k(\mu)\) Delta = get_estimators_from_sim_data(rb_sim_data); This is a scalar computed from the estimates \(\eta^k(\mu)\). Usually the maximum over \(k=0,\ldots,K\) is returned. |
Implements IReducedModel.
Reimplemented in LinEvolDune.ReducedModel.
Definition at line 168 of file ReducedModel.m.
|
static |
Definition at line 157 of file ReducedModel.m.
| function a0 = LinEvol.ReducedModel.rb_init_values | ( | Greedy.DataTree.Detailed.RBLeafNode | detailed_data, |
| decomp_mode | |||
| ) |
function computing initial values for a reduced simulation.
This calls rb_init_values_separable().
| detailed_data | detailed data tree leaf object |
| decomp_mode | flag indicating the operation mode of the function:
|
| a0 | coefficient vector of size N x 1 for the initial values. |
Definition at line 89 of file ReducedModel.m.

| function rb_sim_data = LinEvol.ReducedModel.rb_reconstruction | ( | LinEvol.DetailedData | detailed_data, |
| rb_sim_data | |||
| ) |
(trivial) function computing a detailed reconstruction by linear combination of the coefficients in the simulation data with the orthonormal reduced basis RB
This calls rb_reconstruction_default().
| detailed_data | object defining the basis generation algorithm and storage for storing high dimensional data, i.e. dependent on dimension \(H\). This data is necessary for detailed simulations, construction of online matrices, reduced_data and reconstruction of reduced simulations. |
| rb_sim_data | struct holding reduced simulation data returned by IReducedModel.rb_simulation() . |
| rb_sim_data | struct holding the reduced simulation results and their reconstructions. |
Definition at line 116 of file ReducedModel.m.

| function simulation_data = LinEvol.ReducedModel.rb_simulation_impl | ( | LinEvol.ReducedData | reduced_data | ) |
function, which performs a reduced basis online simulation for the parameter vector \(\mu \in {\cal P} \subset \mathbb{R}^p\), which is assumed to be set by IDetailedModel.set_mu()
The behaviour of this simulation is controlled by the ModelDescr structure rmodel.descr.
if model.name_output_functional is set, then additionally, a sequence of output estimates s(U(:,)) and error bound Delta_s is returned
| reduced_data | an object constructing and storing all (low-dimensional) reduced matrices and vectors needed for reduced simulations. |
| simulation_data | struct holding the reduced simulation data. |
mu_names — the cell array of names of mu-components and for each of these stringt, a corresponding field in model is expected. T — end-time of simulation nt — number of timesteps to compute error_norm — string specifying the used error norm for residual computations. Possible values are l2 or energy. L_I_inv_norm_bound — upper bound on implicit operator inverse norm constant in time, a single scalar L_E_norm_bound — upper bounds on explicit operator constant in time, a single scalar energy_norm_gamma — gamma >= 0 defining the weight in the energy norma0 — a0 LL_E — LL E rb_operators — rb operators LL_I — LL I bb — bb K_II — K II K_IE — K IE K_EE — K EE m_I — m I m_E — m E m — m s_RB — s RB s_l2norm — s l2normdata_const_in_time — if this flag is set, then only operators for first time instance are computed name_output_functional — if this field is existent, then an output estimation is performed and error.estimtations starting_time_step — in t-partition case this is the starting time step for the actual t-partitioni time step stopping_time_step — in t-partition this is the stopping time step for the actual t-partitionDelta0 — initial error added to the total errora — time sequence of solution coefficients, columns are a(:,k) = \(a^{k-1}\) Delta — time sequence of \(L^2\)-posteriori error estimates Delta(k) = \(\Delta^{k-1}_N\) or energy-norm-posteriori error estimates Delta_energy(k) = \(\bar \Delta^{k-1}_N\) depending on the field error_norm in descr. Delta_s — (optional) error bounds for the sequence of output estimates s(U(:,)). This is returned if model.name_output_functional is set. s — (optional) sequence of output estimates s(U(:,)). This is returned if model.name_output_functional is set. LL_I — \(Q_I\) reduced matrix for implicit operator \({\cal L}_{h,I}\) LL_E — \(Q_E\)-sequence of reduced matrices for explicit operator \({\cal L}_{h,E}\) mu — parameter vector of simulation solution Definition at line 19 of file rb_simulation_impl.m.
|
static |
This constant is used for a consistency check of the model descr with help of IModel.struct_check()
Default: struct(" \ 'operators_ptr', {{@(x) isequal(class(x), 'function_handle')}}, \ 'init_values_algorithm', {{@(x) isequal(class(x), 'function_handle')}}, \ 'error_norm', {{@ischar, 'values', {'l2', 'energy'} }} ")
Definition at line 39 of file ReducedModel.m.
 1.8.8
1.8.8